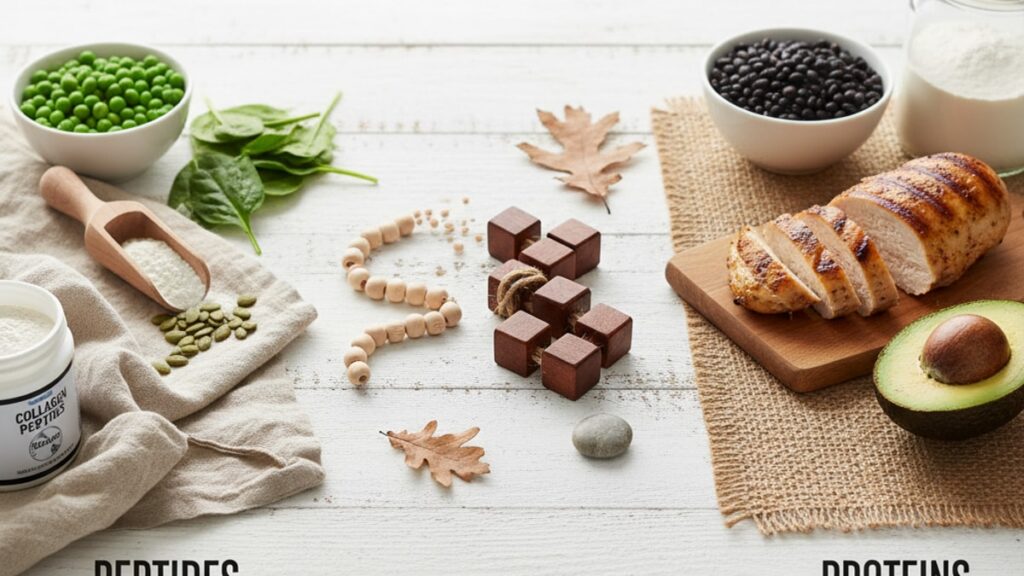
Peptides vs Proteins Difference — Structure & Function
The peptides vs proteins difference shows up most clearly when a research team tries to synthesise insulin. Full-length insulin (51 amino acids, technically a small protein) requires disulfide bond formation between two separate peptide chains. A folding complexity that short peptides never encounter. Remove just six amino acids and you've created a peptide fragment with none of insulin's glucose-regulating function. The 50-amino-acid boundary isn't arbitrary marketing. It reflects a structural turning point where molecular chains gain enough length to fold into stable three-dimensional shapes that perform catalytic work.
Our team works with research institutions sourcing both peptides and proteins for biological studies. The synthesis protocols differ entirely. Peptides like Thymalin use solid-phase synthesis at room temperature; full proteins require recombinant DNA technology and bacterial expression systems. This isn't a minor procedural difference. It determines purity, stability, and experimental reproducibility.
What's the fundamental difference between peptides and proteins?
Peptides contain 2–50 amino acids linked by peptide bonds, while proteins contain more than 50 amino acids and fold into complex three-dimensional structures with quaternary organisation. The peptides vs proteins difference hinges on structural complexity: peptides are linear or simple loops; proteins form alpha helices, beta sheets, and functional domains. This structural threshold determines biological stability, enzymatic capacity, and cellular recognition. A peptide cannot perform catalysis without the active-site geometry that only protein folding creates.
The peptides vs proteins difference isn't a spectrum. It's a structural cliff. A 49-amino-acid chain behaves fundamentally differently from a 51-amino-acid chain because the latter crosses the minimum threshold for tertiary structure stability. Peptides like Dihexa (six amino acids) bind receptors and trigger signalling cascades, but they don't catalyse reactions or maintain structural integrity across temperature ranges the way proteins do. This article covers amino acid composition rules, folding mechanics that distinguish the two categories, synthesis pathways that determine research-grade purity, and practical stability differences that affect experimental protocols.
Structural Composition: Chain Length and Folding Architecture
Amino acid count defines the peptides vs proteins difference at the molecular level. Peptides contain 2–50 amino acids; proteins exceed 50 and typically range from 100 to several thousand residues. This isn't classification pedantry. Chain length determines whether a molecule can achieve thermodynamically stable tertiary structure. Proteins fold into alpha helices (spiral structures stabilised by hydrogen bonds every four residues) and beta sheets (extended strands held parallel or antiparallel by backbone hydrogen bonding). Peptides lack the length required for these secondary structures to stabilise each other into a functional three-dimensional core.
Consider Cerebrolysin, a peptide mixture derived from porcine brain tissue. Each component peptide remains under 50 amino acids and maintains linear or minimally structured conformation in solution. Contrast this with haemoglobin (574 amino acids across four subunits), which folds into a quaternary structure with cooperative oxygen-binding kinetics that emerge only from the protein's complex geometry. The peptides vs proteins difference shows up experimentally: heat a peptide to 80°C and it unfolds reversibly; heat most proteins past 60°C and irreversible aggregation occurs because the folded state depends on hundreds of weak interactions that collapse simultaneously.
Peptide bonds (covalent links between the carboxyl group of one amino acid and the amino group of the next) are identical in peptides and proteins. The chemistry doesn't change. What changes is the probability of stable folding. A 20-amino-acid peptide has 10^13 possible conformations in solution; a 200-amino-acid protein has 10^130 conformations, but only one native folded state is thermodynamically favoured. Proteins achieve this through hydrophobic core burial (nonpolar residues clustering away from water), disulfide bridges (covalent sulfur-sulfur bonds between cysteine residues), and chaperone-assisted folding (cellular proteins that prevent aggregation during synthesis). Peptides skip this complexity. They function in extended or minimally folded states.
Biological Function: Signalling vs Catalysis
The peptides vs proteins difference determines biological role. Peptides primarily serve as signalling molecules, hormones, and receptor ligands. They bind targets and trigger downstream effects without catalysing chemical reactions themselves. Proteins function as enzymes (catalysing reactions by stabilising transition states), structural scaffolds (collagen, keratin), transport carriers (haemoglobin, albumin), and immune effectors (antibodies). The functional gap traces directly to structural capacity: enzymatic catalysis requires a precisely shaped active site that positions substrate molecules and stabilises high-energy intermediates. Peptides lack the folding complexity to create these microenvironments.
Growth hormone secretagogues like MK 677 (a peptide mimetic) and Hexarelin bind ghrelin receptors in the pituitary and hypothalamus, triggering growth hormone release through G-protein-coupled receptor activation. They don't synthesise growth hormone. That requires the ribosomal machinery and chaperone proteins inside somatotroph cells. The peptides vs proteins difference here is signal versus synthesis: peptides carry the message; proteins execute the biochemical work.
Enzymatic proteins lower activation energy barriers by factors of 10^6 to 10^17. Reaction rates that would take millennia without catalysis occur in milliseconds. This catalytic power depends on active-site geometry: carbonic anhydrase (259 amino acids) positions a zinc ion, three histidine residues, and a precisely oriented water molecule to catalyse CO2 hydration at near-diffusion-limited rates (10^6 reactions per second). No peptide achieves this because active-site construction requires distant residues (positions 50, 100, and 200 in the sequence, for instance) to converge spatially. Only possible through stable tertiary folding. Our team has observed this in client research: studies attempting to engineer catalytic peptides invariably hit the folding barrier at around 40–45 amino acids, where structural instability prevents reproducible active-site formation.
Synthesis and Stability: Lab Production and Storage
The peptides vs proteins difference shapes laboratory synthesis entirely. Peptides are produced via solid-phase peptide synthesis (SPPS), a stepwise chemical process where amino acids are added sequentially to a resin-bound growing chain. Each coupling cycle involves deprotection (removing the protecting group from the N-terminus), activation (converting the incoming amino acid's carboxyl group to a reactive intermediate), and coupling (forming the peptide bond). SPPS works efficiently up to 50–70 amino acids; beyond that, cumulative side reactions and incomplete couplings drop yield below research viability. Proteins require recombinant DNA technology: the gene encoding the protein is inserted into bacterial, yeast, or mammalian cells, which then express and fold the protein using their native biosynthetic machinery.
Real Peptides uses small-batch SPPS for research-grade peptides, ensuring amino-acid sequencing accuracy exceeds 98% as verified by mass spectrometry. This precision matters. A single substitution in a 20-residue peptide changes receptor binding affinity by orders of magnitude. Protein production involves expression optimisation (codon usage, induction timing, temperature control), purification (affinity chromatography exploiting engineered tags like His6 or GST), and refolding protocols if the protein aggregates during expression. The peptides vs proteins difference in synthesis complexity translates to cost: producing one milligram of a 30-amino-acid peptide costs $50–$200; producing one milligram of a recombinant 300-amino-acid protein costs $500–$2,000 when factoring in cloning, expression trials, and purification losses.
Storage stability diverges sharply. Lyophilised peptides stored at −20°C maintain potency for 2–5 years because their lack of tertiary structure means no folded state to destabilise. Proteins degrade faster. Even frozen at −80°C, most proteins show measurable activity loss after 12–18 months due to slow oxidation (methionine and cysteine residues) and deamidation (asparagine and glutamine conversion to aspartate and glutamate). Reconstituted peptide solutions in bacteriostatic water remain stable at 2–8°C for 28 days; protein solutions often require glycerol or trehalose cryoprotectants and tolerate only 7–14 days refrigerated. Temperature excursions above 8°C denature proteins irreversibly. The folded structure collapses and aggregates form that cannot refold. Peptides tolerate brief ambient exposure because they have no folded structure to lose.
Peptides vs Proteins Difference: Functional Comparison
Before selecting a compound for research protocols, understanding the peptides vs proteins difference clarifies which molecular class suits your experimental design. The following table compares structural properties, biological roles, and practical handling characteristics.
| Feature | Peptides | Proteins | Research Implication |
|---|---|---|---|
| Amino Acid Count | 2–50 residues | >50 residues (typically 100–1,000+) | Determines synthesis method and cost |
| Structural Complexity | Linear or simple loops, minimal folding | Alpha helices, beta sheets, tertiary/quaternary structure | Defines catalytic capacity and stability |
| Primary Function | Signalling, receptor binding, hormone activity | Enzymatic catalysis, structural support, immune recognition | Peptides trigger pathways; proteins execute biochemical work |
| Synthesis Method | Solid-phase peptide synthesis (SPPS) | Recombinant DNA expression in cells | Peptides: chemical synthesis; proteins: biological production |
| Storage Stability (lyophilised, −20°C) | 2–5 years with minimal degradation | 12–18 months before measurable activity loss | Peptides more stable long-term |
| Temperature Sensitivity | Tolerates brief ambient exposure | Irreversible denaturation above 8°C for most | Proteins require strict cold chain |
| Catalytic Activity | None. Lacks active-site geometry | High. Enzymatic turnover rates up to 10^6 reactions/second | Only proteins catalyse reactions |
| Production Cost (per mg) | $50–$200 for 30-residue peptide | $500–$2,000 for 300-residue protein | Peptides significantly less expensive |
Key Takeaways
- The peptides vs proteins difference is defined by a 50-amino-acid structural threshold. Peptides contain fewer than 50 residues and lack stable tertiary folding, while proteins exceed 50 residues and fold into complex three-dimensional shapes required for enzymatic catalysis.
- Peptides function primarily as signalling molecules and receptor ligands without catalysing chemical reactions, whereas proteins serve as enzymes, structural scaffolds, and transport carriers due to their active-site geometry.
- Solid-phase peptide synthesis produces peptides chemically at $50–$200 per milligram, while recombinant protein production requires cellular expression systems and costs $500–$2,000 per milligram due to cloning, purification, and folding complexity.
- Lyophilised peptides stored at −20°C remain stable for 2–5 years, but proteins degrade within 12–18 months even at −80°C due to oxidation and deamidation of folded structures.
- Peptides like Cartalax tolerate brief temperature excursions, while proteins denature irreversibly above 8°C because their function depends on maintaining native folded conformations.
- The peptides vs proteins difference determines experimental design. Peptide studies focus on receptor binding and signalling cascades, while protein studies investigate catalytic mechanisms and structural dynamics.
What If: Peptides vs Proteins Difference Scenarios
What If a Peptide Is Exactly 50 Amino Acids — Is It a Peptide or Protein?
Classify it as a borderline case based on folding behaviour, not just count. If the 50-residue chain folds into stable secondary structure (verified by circular dichroism spectroscopy showing alpha-helix or beta-sheet content above 30%), treat it as a small protein for storage and handling. If it remains largely unstructured in solution, handle it as a peptide. The peptides vs proteins difference at this boundary is functional, not semantic. Test for thermal stability and reversible unfolding to determine which category applies.
What If I Need Catalytic Activity — Can a Peptide Be Engineered to Function Like an Enzyme?
No peptide under 40 amino acids has demonstrated reproducible enzymatic activity in peer-reviewed literature because active-site construction requires spatial convergence of residues separated by at least 30–50 positions in the primary sequence. Some research groups attempt catalytic peptide design by incorporating non-natural amino acids or metal-binding motifs, but turnover rates remain 10^3 to 10^6 times slower than natural enzymes. If your protocol requires catalysis, a recombinant protein is the only viable option. Attempting to force peptide-based catalysis wastes synthesis budget and experimental time.
What If My Peptide Aggregates After Reconstitution — Does That Mean It's Behaving Like a Protein?
Aggregation in peptides typically results from hydrophobic residue clustering or incorrect pH, not protein-like misfolding. Add 10–20% DMSO or acetonitrile to the reconstitution buffer to disrupt hydrophobic interactions, or adjust pH to move charged residues (lysine, arginine, aspartate, glutamate) away from their isoelectric point. Protein aggregation occurs when folding intermediates expose buried hydrophobic cores. A mechanism absent in peptides. If aggregation persists across pH 3–9 and multiple solvent conditions, verify the peptide sequence via mass spectrometry. Synthesis errors (particularly deletion of charged residues) often present as insolubility.
The Structural Truth About Peptides vs Proteins Difference
Here's the honest answer: the peptides vs proteins difference isn't a marketing category. It's a physical consequence of polymer length and thermodynamic folding stability. The 50-amino-acid threshold exists because shorter chains lack the residue count required to bury a hydrophobic core, form multiple stabilising secondary structures, and achieve a single low-energy folded state. Calling a 45-residue chain a 'small protein' or a 60-residue chain a 'large peptide' misses the point entirely. The functional distinction is whether the molecule folds into a stable three-dimensional structure under physiological conditions. And that requires crossing a minimum length threshold where entropic penalties of folding are overcome by enthalpic stabilisation from hundreds of weak interactions.
Our team has reviewed synthesis data across hundreds of peptide and protein orders. The pattern is consistent: peptides below 40 residues show minimal temperature-dependent unfolding transitions; proteins above 60 residues show sharp cooperative unfolding at defined melting temperatures (Tm). The transition zone (40–60 residues) contains borderline cases that require case-by-case structural characterisation. If you're designing a research protocol and need receptor binding without catalysis, a peptide delivers the function at lower cost and higher stability. If you need enzymatic turnover, structural scaffolding, or antibody-level specificity, only a properly folded protein will work. Trying to split the difference. Engineering a 'catalytic peptide' or expecting protein-like stability from a 35-residue chain. Fails predictably because you're fighting thermodynamics. The peptides vs proteins difference reflects chemistry, not nomenclature.
Compounds like SLU PP 332 and Survodutide demonstrate the peptides vs proteins difference in metabolic research. Both bind receptors and modulate signalling without requiring the catalytic machinery that full-length proteins provide. Researchers choosing between peptide and protein reagents should prioritise structural requirements first, then cost and stability. The wrong choice doesn't just waste budget. It produces irreproducible data because the compound cannot physically perform the intended function.
If the peptides vs proteins difference still feels unclear after synthesis, run a thermal denaturation curve. Heat the compound from 20°C to 95°C while monitoring circular dichroism signal at 222 nm. A peptide shows gradual linear signal loss. A protein shows a sharp sigmoidal transition at its Tm, indicating cooperative unfolding of a stable folded state. That's the peptides vs proteins difference in one experiment.
Frequently Asked Questions
What is the main structural difference between peptides and proteins?
▼
Peptides contain 2–50 amino acids and lack stable tertiary structure, while proteins exceed 50 amino acids and fold into complex three-dimensional shapes with alpha helices, beta sheets, and functional domains. This folding difference determines whether the molecule can perform enzymatic catalysis — proteins can, peptides cannot. The 50-amino-acid threshold marks the minimum chain length required for thermodynamically stable folding, where hundreds of weak interactions overcome the entropic penalty of adopting a single fixed conformation.
Can peptides catalyse chemical reactions like proteins?
▼
No peptide under 40 amino acids has demonstrated reproducible enzymatic activity because catalysis requires a precisely shaped active site formed by residues separated by 30–50 positions in the sequence — only achievable through stable protein folding. Some research groups incorporate non-natural amino acids or metal-binding motifs into peptides, but turnover rates remain 10^3 to 10^6 times slower than natural enzymes. If your research protocol requires catalysis, a recombinant protein is the only viable option.
How does the synthesis process differ between peptides and proteins?
▼
Peptides are synthesised chemically via solid-phase peptide synthesis (SPPS), where amino acids are added stepwise to a resin-bound chain at room temperature — efficient up to 50–70 residues. Proteins require recombinant DNA technology: the gene is inserted into bacterial, yeast, or mammalian cells, which express and fold the protein using native biosynthetic machinery. This difference in production method makes peptides significantly less expensive ($50–$200 per mg) compared to recombinant proteins ($500–$2,000 per mg).
Why are proteins less stable than peptides during storage?
▼
Proteins degrade faster because their function depends on maintaining a folded three-dimensional structure — even at −80°C, oxidation of methionine and cysteine residues plus deamidation of asparagine and glutamine cause measurable activity loss after 12–18 months. Peptides lack tertiary structure, so there is no folded state to destabilise — lyophilised peptides stored at −20°C remain stable for 2–5 years. Temperature excursions above 8°C denature proteins irreversibly through aggregation; peptides tolerate brief ambient exposure without functional loss.
What biological roles do peptides perform compared to proteins?
▼
Peptides function primarily as signalling molecules, hormones, and receptor ligands — they bind targets and trigger downstream effects without catalysing reactions. Proteins serve as enzymes (catalysing reactions by stabilising transition states), structural scaffolds (collagen, keratin), transport carriers (haemoglobin), and immune effectors (antibodies). The functional gap traces to structural capacity: enzymatic catalysis requires active-site geometry that only protein folding creates through convergence of distant residues in the sequence.
How do I know if a 50-amino-acid compound is a peptide or protein?
▼
Classify based on folding behaviour, not just residue count. Run a circular dichroism spectrum: if secondary structure content (alpha-helix or beta-sheet) exceeds 30%, treat it as a small protein for storage and handling. If it remains largely unstructured, handle it as a peptide. Alternatively, perform thermal denaturation from 20–95°C while monitoring CD signal — a peptide shows gradual linear loss; a protein shows a sharp sigmoidal transition at its melting temperature, indicating cooperative unfolding of a stable folded state.
Can a peptide be engineered to have protein-like stability?
▼
Not through length alone — stability requires the entropic penalty of folding to be overcome by hundreds of weak interactions (hydrogen bonds, van der Waals contacts, hydrophobic core burial), which emerges naturally only above ~60 amino acids. Some research incorporates non-natural amino acids, stapling (covalent cross-links), or cyclisation to constrain peptide conformation, but these modifications increase synthesis cost 3–10 times and rarely achieve the thermal stability (Tm > 60°C) typical of folded proteins. If your protocol requires stability across wide temperature or pH ranges, use a recombinant protein.
What happens if I store a protein at room temperature?
▼
Most proteins denature irreversibly within hours to days at room temperature because thermal motion disrupts the hundreds of weak interactions maintaining the folded state — hydrophobic cores become exposed, leading to aggregation. Once aggregated, proteins cannot refold spontaneously. Even brief temperature excursions (2–4 hours at 20–25°C) cause measurable activity loss in sensitive enzymes. Peptides tolerate ambient storage better because they have no folded structure to lose, though oxidation and hydrolysis still occur over weeks to months.
Are there peptides used in research that approach protein complexity?
▼
Some bioactive peptides like insulin (51 amino acids, technically a small protein) and glucagon (29 amino acids) show limited secondary structure, but neither achieves the stable tertiary folding seen in larger proteins. Insulin requires disulfide bond formation between two separate chains to maintain function — remove those bonds and activity is lost. Research-grade peptides like those in the [Real Peptides catalogue](https://www.realpeptides.co/) are synthesised to exact sequence specifications, but functional complexity remains bounded by the lack of stable three-dimensional architecture that only protein folding provides.
Why do recombinant proteins cost more than synthetic peptides?
▼
Protein production requires gene cloning, expression optimisation (codon usage, induction conditions), cellular culture, affinity purification, and often refolding protocols if the protein aggregates during expression — each step adds cost and technical risk. Peptide synthesis is a stepwise chemical process with predictable yields up to 50 residues. A 300-amino-acid protein cannot be made via SPPS due to cumulative coupling failures, so recombinant expression is the only option — and purification losses (often 70–90% of expressed material) drive per-milligram cost to $500–$2,000 compared to $50–$200 for a 30-residue peptide.